AMHERST, Mass. – Studies led by cell biologist Thomas Maresca at the University of Massachusetts Amherst are revealing new details about a molecular surveillance system that helps detect and correct errors in cell division that can lead to cell death or human diseases. Findings are reported in the current issue of the Journal of Cell Biology.
The purpose of cell division is to evenly distribute the genome between two daughter cells. To achieve this, every chromosome must properly interact with a football-shaped structure called the spindle. However, interaction errors between the chromosomes and spindle during division are amazingly common, occurring in 86 to 90 percent of chromosomes, says Maresca, an expert in mitosis.
“This is not quite so surprising when you realize that every single one of the 46 chromosomes has to get into perfect position every time a cell divides,” he notes. The key to flawless cell division is to correct dangerous interactions before the cell splits in two.
Working with fruit fly tissue culture cells, Maresca and graduate students Stuart Cane and Anna Ye have developed a way to watch and record images of the key players in cell division including microtubule filaments that form the mitotic spindle and sites called kinetochores that mediate chromosome-microtubule interactions. They also examined the contribution of a force generated by molecular engines called the polar ejection force (PEF), which is thought to help line up the chromosomes in the middle of the spindle for division. For the first time, they directly tested and quantified how PEF, in particular, influences tension at kinetochores and affects error correction in mitosis.
“We also now have a powerful new assay to get at how this tension regulates kinetochore-microtubule interactions,” Maresca adds. “We knew forces and tension regulated this process, but we didn’t understand exactly how. With the new technique, we can start to dissect out how tension modulates error correction to repair the many erroneous attachment intermediates that form during division.”
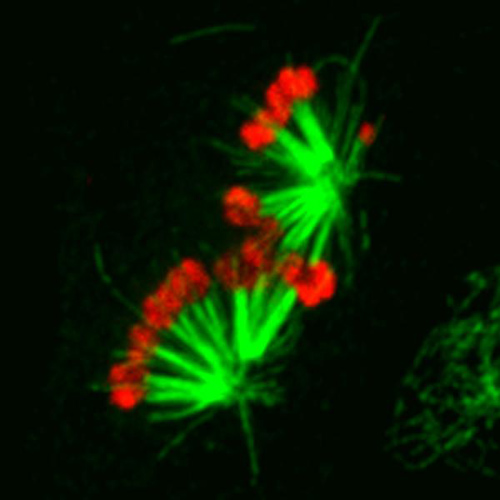
Expression of the chromokinesin NOD (red) stabilizes aberrant interactions between kinetochores and spindle microtubules (green). Tension stabilizes bioriented attachments where each sister chromatid is attached to microtubules from opposite spindle poles while tensionless attachments are typically unstable and corrected. Elevating the polar ejection force that pushes chromosome arms away from spindle poles overwhelms error correction, resulting in a dose-dependent stabilization of syntelic attachments where sister chromatids are attached to the same spindle pole. Image by Tom Maresca
The cell biologists conduct their experiments inside living cells. In normal cell division, chromosomes line up in the center, where two copies of each chromosome are held together with “molecular glue” until signaled to dissolve the glue and divide. To oversimplify, each chromosome copy is then pulled to opposite poles of the cell, escorted in what looks like a taffy pull away from the center as two new daughter cells are formed.
During the split, molecular engines pull the copies apart along microtubule tracks that take an active role in the process that includes shortening microtubules by large, flexible scaffold-like protein structures called kinetochores that assemble on every chromosome during division. Maresca and colleagues say until this study revealed details, PEF’s function as a kinetochore regulator has been underappreciated.
Overall, this well-orchestrated process prevents serious problems such as aneuploidy, that is, too many chromosomes in daughter cells. Aneuploidy in somatic or body cells leads to cell death and is a hallmark of most cancer cells. But in eggs or sperm, it leads to serious birth defects and miscarriages.
In properly aligned division, microtubules from opposite spindle poles tug chromosome copies toward opposite poles, but they stick together with molecular glue until the proper moment. This creates tension at the kinetochores and stabilizes their interactions with microtubules. However, if attachments are bad, or syntelic, both copies attach to the same pole, leading to chromosome mis-segregation and aneuploidy if uncorrected. “Cells have a surveillance mechanism that allows them to wait for every chromosome to properly align before divvying up the chromosomes,” Maresca says. “It’s clear in our movies that the cell waits for the last kinetochores to correctly orient before moving forward.”
To study this at the molecular level, Maresca and colleagues designed experiments to trick the cellular machinery into overexpressing the molecular engine that produces PEF. Surprisingly, this caused a dramatic increase in a type of bad kinetochore-microtubule interaction called syntelic attachments. They also fluorescently tag chromosomes, microtubules, kinetochores and the molecular engine kinesin with different colors to visualize interactions in real time using a special microscope at UMass Amherst able to image single molecules. Quantifying the amount of fluorescence of the force-producing molecular engine, they were able to assess the relative strength of the PEF in cells.
Maresca says, “We see the detection pathway preventing the molecular glue from dissolving until every chromosome is correctly aligned. The delay gives the cell time to correct errors. We propose that these bad syntelic attachments are normally very short-lived because they are not under proper tension. However, when we experimentally elevate PEF, tension is introduced at attachments that do not typically come under tension, essentially tricking the cell into thinking these chromosomes are properly aligned.”
Plotting the percent of syntelic attachments versus the amount of PEF, Maresca and colleagues observed an error rate that plateaus at 80 to 90 percent, mirroring and supporting earlier work by others in different cell types. “In cells with elevated PEFs, the correction pathway is overridden, the detection mechanism is silenced and the result is disastrous because it leads to severe aneuploidy. This research has taught us about how an important molecular engine generates the PEF and how this force affects the accuracy of cell division.”
*Source: University of Massachusetts Amherst
