Masonic Cancer Center researchers discover a virus-fighting enzyme
It’s well known that sunlight and chemical carcinogens can mutate DNA, and that mutations are essential for cancer to develop.
One big mystery was what causes the thousands of mutations evident in most breast cancers.
Now researcher Reuben Harris, Masonic Cancer Center, University of Minnesota, and his colleagues have found evidence that one of our own proteins is a major source of these mutations. The researchers have just published evidence implicating the protein—an enzyme called APOBEC3B—in the international journal Nature.

Reuben Harris led a team that discovered a major source of DNA mutations in breast cancer. Image credit: University of Minnesota
“We’ve discovered a rogue DNA-mutating enzyme in more than half of breast cancers. This is the first clear example of enzyme-induced mutations in human cancer,” says Harris, an associate professor in the Department of Biochemistry, Molecular Biology & Biophysics. “With this knowledge, we might be able to do something about it.
“An obvious next step is to determine how early APOBEC3B starts contributing to mutation in breast cancer, and figuring out ways to suppress the accumulation of these mutations. This could have a major impact—for instance, in decreasing the chance of tumors developing resistance to current therapeutics.”
Calling card for cancer?
APOBEC3B belongs to a family of enzymes called APOBECs (A’-po-becks). They function, earlier work in Harris’s lab found, to mutate DNA. Some, active in white blood cells, mutate HIV-derived DNA as a first line of defense against the virus. APOBEC3B also seems to act normally against viruses and other parasitic elements.
But Harris and his colleagues turned up several pieces of evidence pointing to APOBEC3B as a source of mutations in breast cancer. For example, they discovered elevated APOBEC3B levels in many breast cancer cell lines and fresh breast tumor cells (but not normal breast cells) they studied. And disabling the gene for APOBEC3B in breast cancer cell lines slashed the number of mutations and established clear cause and effect.
They further showed that APOBEC3B tends to mutate DNA in a certain pattern, which is written all over most breast cancer genomes and is equivalent to a “calling card.”
Gutting a guardian
Consider one of the best-known “cancer genes,” TP53, which functions to suppress cancer and is often referred to in scientific circles as “the guardian of the genome.” Working with large data sets, the researchers found it is mutated—and inactivated—in breast tumors with high APOBEC3B activity.
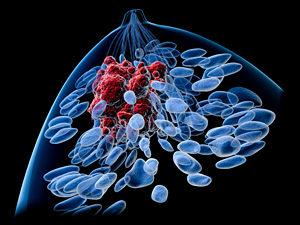
Top view of a female breast with a tumor. Features body silhouette, lobules, lactiferous ducts and tumor. Image credit: University of Minnesota
The best explanation for this, says Harris, is that something turns on APOBEC3B activity in cells that are otherwise healthy. It then mutates the TP53 gene and inactivates it, unleashing a chain of events that results in cancer.
“We showed that APOBEC3B causes chromosomal damage and mutagenesis,” Harris explains. “Also, we see higher APOBEC3B activity at all stages of breast cancer, implying it’s an early event. So we favor APOBEC3B as the cause of TP53 inactivation.”
Inessential gene, enigmatic enzyme
“APOBEC3B is a double-edged sword,” says Harris. “It’s a really good [antiviral] enzyme, but its DNA-mutating activity can also be detrimental to the cell. Obviously, this has been balanced by evolution, and it generally doesn’t become a problem until near the end of reproductive age.”
But it isn’t essential for life; although present in many Europeans and residents of sub-Saharan Africa, the gene for APOBEC3B is lacking in many Pacific Islanders and Asians, notably Japanese, who have a very low rate of breast cancer.
Harris would like to know how the absence or presence of APOBEC3B and its gene correlates with cancer risk and other factors.
“We are confident that our breakthrough will trigger a domino effect of additional research and clinical studies that result in better outcomes for patients with breast cancer,” he says.
*Source: University of Minnesota
