Joint BioEnergy Institute Researchers Combine Systems Biology with Genetic Engineering to Improve Production of Isopentenol in E.Coli
In the on-going effort to develop advanced biofuels as a clean, green and sustainable source of liquid transportation fuels, researchers at the U.S. Department of Energy (DOE)’s Joint BioEnergy Institute (JBEI) have identified microbial genes that can improve both the tolerance and the production of biogasoline in engineered strains of Escherichia coli.
Aindrila Mukhopadhyay, a chemist who directs the host engineering program for JBEI’s Fuels Synthesis Division, led a study in which transcriptomic data and a synthetic metabolic pathway were used to identify several genes that not only improve tolerance but also production of isopentenol in E.coli. Isopentenol is a five-carbon alcohol that is a highly promising candidate for biogasoline, but, like other short-chained alcohols, is toxic to E.coli at commercial levels of fuel production.
“Our study demonstrates that microbial tolerance engineering using transcriptomics data can be used to identify target genes that improve fuel production,” says Mukhopadhyay, who also holds an appointment with the Lawrence Berkeley National Laboratory (Berkeley Lab)’s Physical Biosciences Division. “Our targets include a regulator for amino acid biosynthesis, and an ABC transporter protein, the first native transporter that improves tolerance to a short-chain alcohol.”

Aindrila Mukhopadhyay (left) and Heather Jensen were part of a JBEI team that identified microbial genes which can improve both the tolerance and the production of biogasoline in engineered strains of E. coli. (Photo by Roy Kaltschmidt)
Mukhopadhyay is the corresponding author of a paper describing this study in the journal mBio titled “Improving microbial bio-gasoline production in Escherichia coli using tolerance engineering.” Co-authors are Heather Jensen, Jee Loon Foo, Robert Dahl, Kevin George, Jay Keasling, Taek Soon Lee and Susanna Leong.
The price of gasoline and other petroleum fuels may be dropping for the moment, but atmospheric carbon concentrations are continuing to rise. A highly touted carbon-neutral alternative to petroleum fuels is the microbial production of advanced biofuels from the cellulosic biomass of perennial grasses and other non-food plants, as well as from agricultural waste. However, the toxicity to microbes of many of the best candidate compounds for advanced biofuels presents a “production versus survival” conundrum.
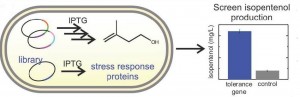
The overexpression of E. coli genes displaying tolerance to isopentenol increased production of this leading biogasoline candidate in a production strain of the microbes. Image credit: Berkeley Lab (Click image to enlarge)
“In order for microbial biofuel production to be cost effective, yields must exceed native microbial tolerance levels, necessitating the development of solvent-tolerant microbial strains,” Mukhopadhyay says. “In parallel with improved tolerance it is also crucial that we improve production.”
To this end, Mukhopadhyay and her group in this new study used transcriptomic data – a measurement of differential expression of gene transcripts in a given genome – to identify 40 E.coli genes that showed increase when exposed to externally added isopentenol. These genes were then overexpressed in E. coli to evaluate their potential for improving isopentenol tolerance. Genes conferring isopentenol tolerance were then co-expressed individually with an isopentenol production metabolic pathway in E.coli to determine which would increase productivity as well.
“MetR, the methionine biosynthesis regulator, improved the titer for isopentenol production by 55-percent,” Mukhopadhyay says. “MdlB, the ABC transporter, facilitated a 12-percent improvement in isopentenol production.”
Mukhopadhyay and her group are especially eager to further investigate the MdlB transporter, which they believe, as the first native transporter gene shown to improve production of a short-chain alcohol, will provide a valuable new avenue for host engineering in biogasoline production.
“The critical point is that you must first identify the genes that can serve as engineering targets, and then test them to find which ones work best,” Mukhopadhyay says. “Now that we have identified MdlB as a target, we are going to examine it in great depth to see how can we improve its function and optimize its use in a production microbe.”
This research was supported by the DOE Office of Science. JBEI is a DOE Office of Science Bioenergy Research Center led by Berkeley Lab.
Additional Information
For more about the research of Aindrila Mukhopadhyay go here
For more about JBEI go here
*Source: Berkeley Lab
